
TOKYO, Sept. 9, 2022 /PRNewswire/ — In a study published recently in the journal Nature, researchers from Keio University and colleagues identify genes involved in the development of severe COVID-19 in the younger Japanese population.
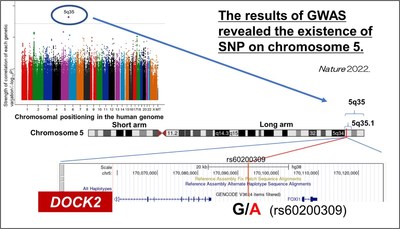
Genes play a key role in the development and progression of infectious diseases. While studies have identified high-risk genes for COVID-19 in Europeans, there is limited genetic data for the Japanese population. Now, an inter-Japan research collaboration has provided critical insights based on detailed genetic analysis. Koichi Fukunaga from Keio University and his colleagues show that a gene important for immune function is involved in severe COVID-19, especially in the younger Japanese population.
For their investigation, the researchers compared the genomes of 2,393 patients who suffered from COVID-19 with that of 3,289 healthy individuals. All COVID-19 patients had been hospitalized while infected and 990 of them had been afflicted by a severe form of the disease.
The researchers identified seven genes showing prominent differences between the two groups, of which six were more prevalent in younger patients (less than 65 years old) and in the more severe cases. Upon further examination of these populations, the team conducted a genetic analysis of this population revealing that the gene responsible for producing a protein called DOCK2 was mutated in these subjects.
Blood samples of COVID-19 patients were then sequenced to reveal that the risk of a mutation on this gene was associated with low levels of the DOCK2 protein. DOCK2 is currently widely understood to play a role in immune signaling, so to understand its role in the context of COVID-19 progression, the researchers isolated the blood cells of a smaller group of subjects. Their analysis showed particularly low values of DOCK2 in some white blood cells. When this observation was replicated in lab cells, using a drug that inhibits DOCK2, the cells showed hampered movement and an inability to release chemical messengers that warn the body of disease.
The worst region of the body to be affected by COVID-19 is often the lungs, often resulting in pneumonia. Therefore, lung samples of three deceased COVID-19 patients were scrutinized. DOCK2 was found to be suppressed in the cells of these samples. Finally, the team studied the real-time effects of decreased DOCK2 in hamsters infected with SARS-CoV-2. These hamsters showed greater weight loss and fluid buildup in their lungs compared to healthy ones. The migration of immune cells—which is essential in combating the virus—was also compromised in the infected hamsters.
This genome-wide study reveals the potential role of DOCK2 in influencing the severity of COVID-19, especially in younger populations. It could also be used as a target for drug development to fight the infection. “DOCK2 plays a key role in host immune response to SARS-CoV-2 infection and development of severe COVID-19, being a promising biomarker and therapeutic target,”1
Reference
- Namkoong, H. et al. DOCK2 is involved in the host genetics and biology of severe COVID-19. Nature (2022) doi:10.1038/s41586-022-05163-5.
Background to the research in this paper and related information
DOCK2 and the immune system: When the DOCK2 gene is activated in our cells, it passes on the green light for DOCK2 protein synthesis.
The DOCK2 protein is known to play a role in the production and release of interferons, which are chemical messengers released by infected cells to get help from distant immune cells. DOCK2 also helps certain types of white blood cells move and migrate to the site of infection (a process known as chemotaxis) to fight the invading pathogens. These mechanisms are imperative to reducing infection, and why researchers believe younger COVID-19 patients, who are at higher risk for mutations of the DOCK2 gene, showed more severe forms of the disease.
The Joint Research Coronavirus Task Force
The paper described in this press release is the culmination of recent inter-Japan collaborative research conducted by The Joint Research Coronavirus Task Force (Japan COVID-19 Task Force) comprising of more than 120 hospitals across Japan, including 13 facilities affiliated with Keio University.1
The Japan COVID-19 Task Force was launched in May 2020 to offer a genetic approach to analyzing the spread and treatment of COVID-19.
Professor Koichi Fukunaga—lead author of the paper and specialist in respiratory medicine at the Keio University School of Medicine in Tokyo—is the head of the Task Force that has continued to collect clinical information and samples from patients during each virus variant epidemic and collaborated with hospitals throughout Japan.
The Japan COVID-19 Task Force is focusing on trends in Japan, including clarifying why the number of deaths in Japan has been significantly lower than in the United States and Europe. The results of the studies from Japanese patients are being shared with researchers worldwide to contribute to global efforts to develop treatments for COVID-19.
- Interview with Professor Koichi Fukunaga: “Joint Research Coronavirus Task Force: The power of integrating clinical and genetic research to understand and eliminate COVID-19”
Keio Research Highlights: https://research-highlights.keio.ac.jp/2022/01/a.html
Further information
Office of Research Development and Sponsored Projects
Keio University
2-15-45 Mita, Minato-ku, Tokyo 108-8345 Japan
Telephone: +81 (0)-3-5427-1678
E-mail: keio-rpr@adst.keio.ac.jp
Websites
Keio University
https://www.keio.ac.jp/en/
Keio Research Highlights
https://research-highlights.keio.ac.jp/
Photo: https://mma.prnewswire.com/media/1895226/fig_1.jpg
Source link
The content is by PR Newswire. Headlines of Today Media is not responsible for the content provided or any links related to this content. Headlines of Today Media is not responsible for the correctness, topicality or the quality of the content.

